High-Fidelity Equivariant Cryo-Electron Tomography
Vinith Kishore1 , Valentin Debarnot2 , Ricardo D. Righetto3 , Benjamin D. Engel3 , Ivan Dokmanić1 .
1 Department of Mathematics and Computer Science, University of Basel,
2 INSA‐Lyon, Universite Claude Bernard Lyon 1, CNRS, Inserm, CREATIS UMR 5220, U1294,
3 Biozentrum, University of Basel.
Icecream is a self-supervised framework for cryo-ET (and standard ET) reconstruction that integrates equivariance principles from modern imaging theory into a deep-learning architecture. This webpage aims at providing reconstruction example and documentation on how to use the code.
Table of Contents
- What to expect with Icecream?
- How to use Icecream
- Examples of reconstructions
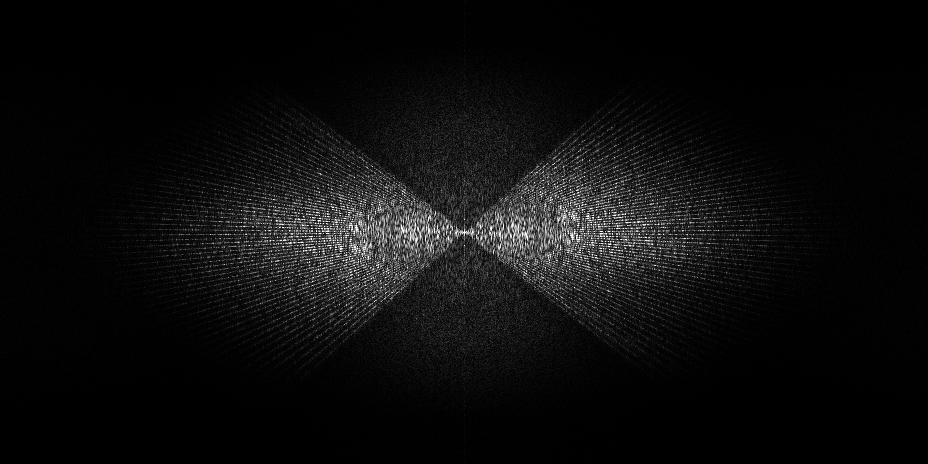
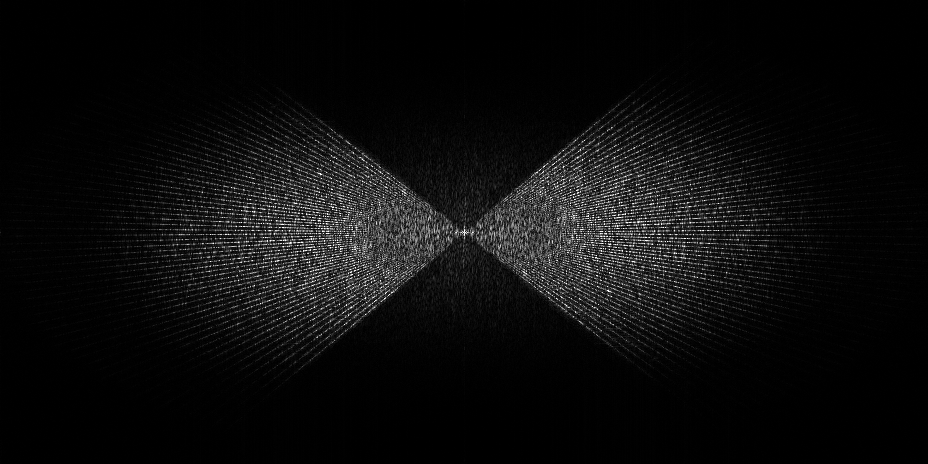
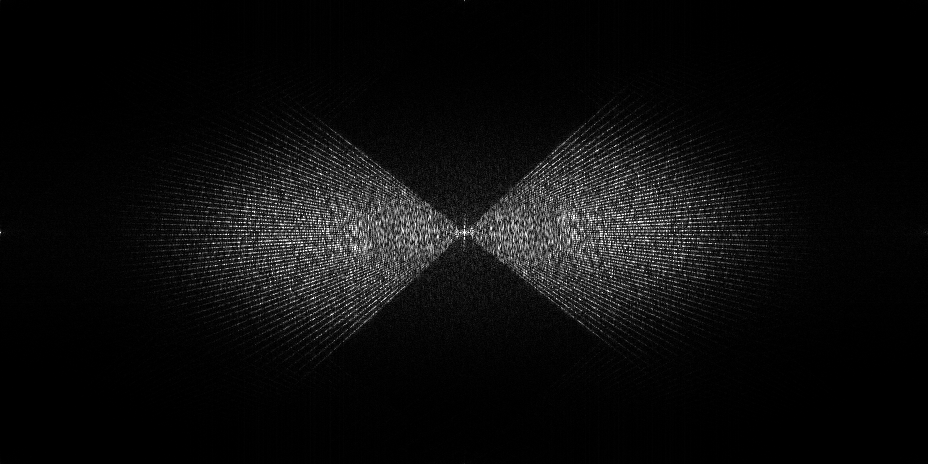
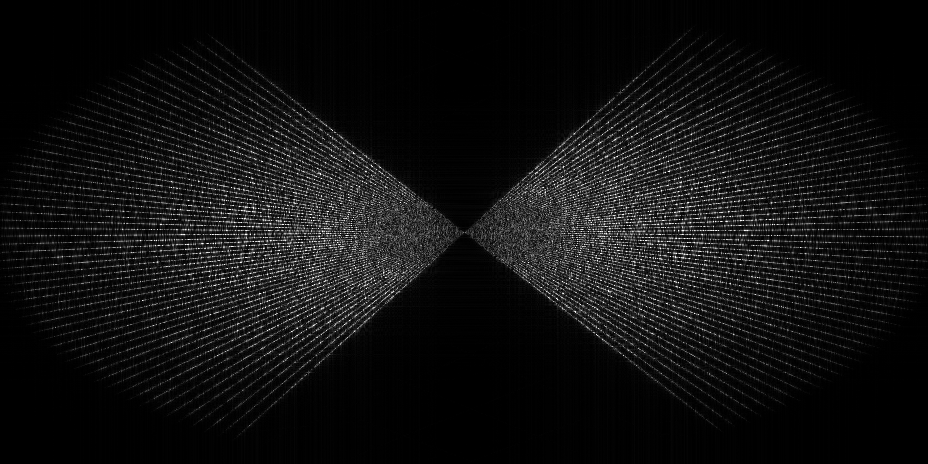
- Dose vs angle splitting
- How to cite
What to expect with Icecream?
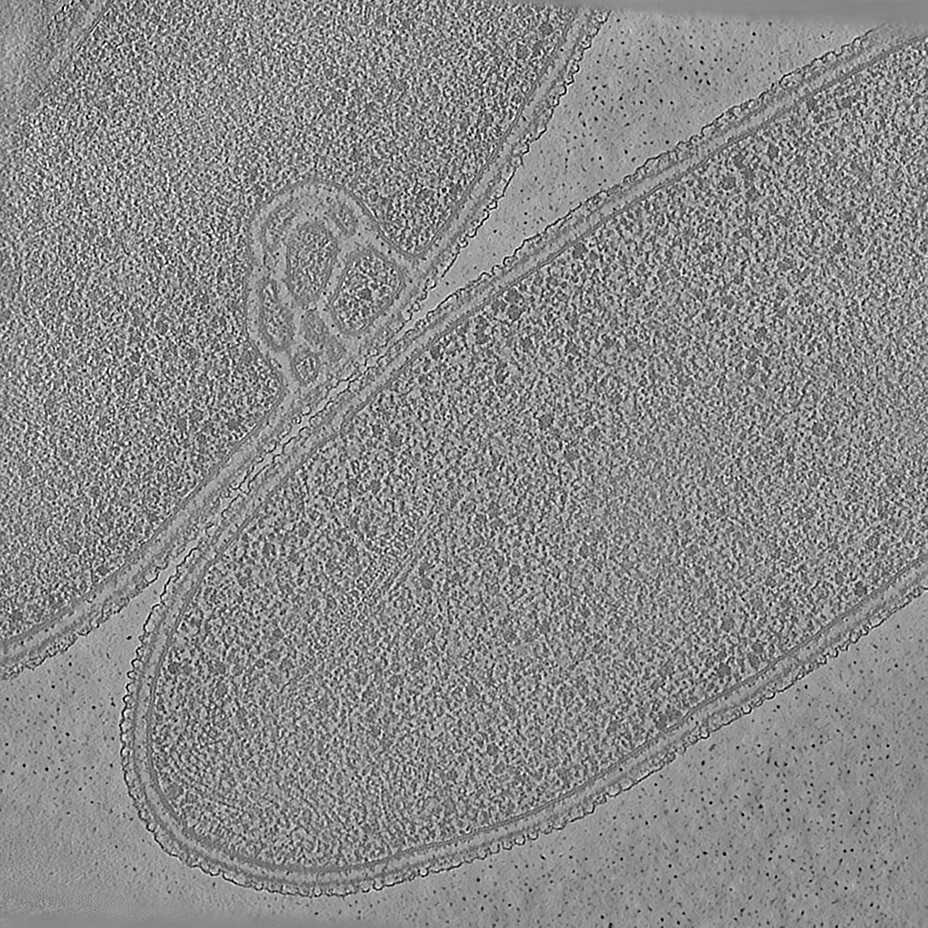
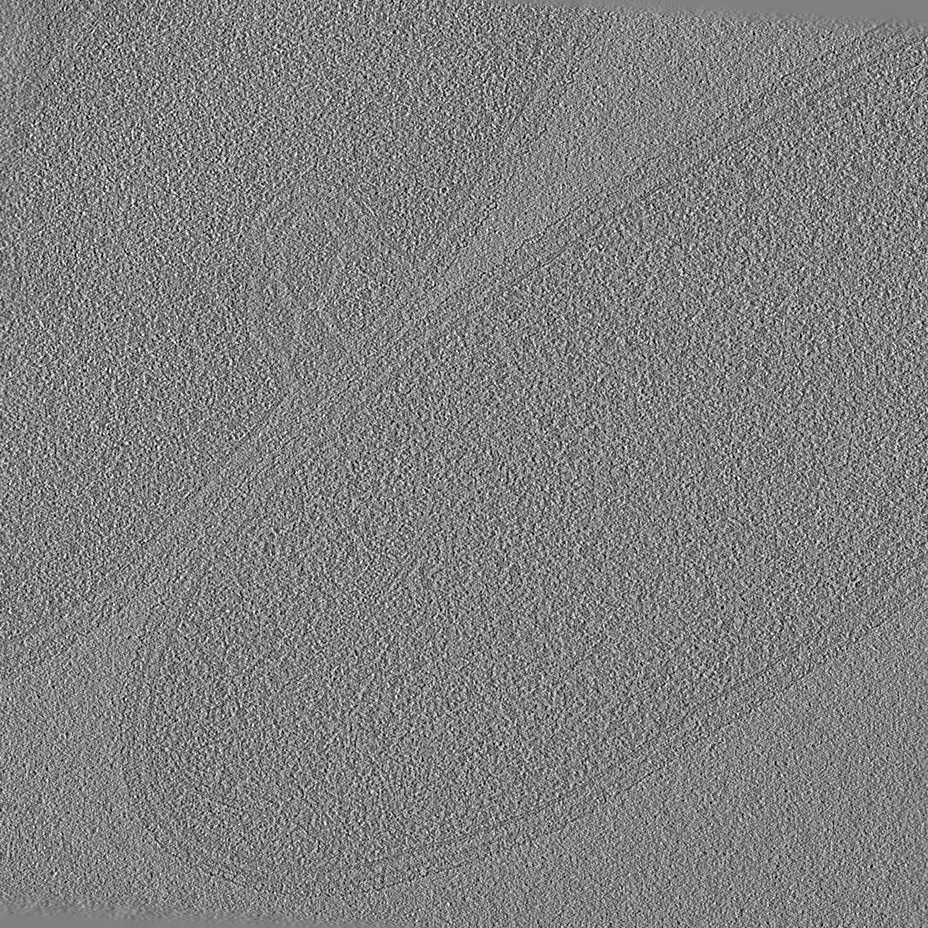
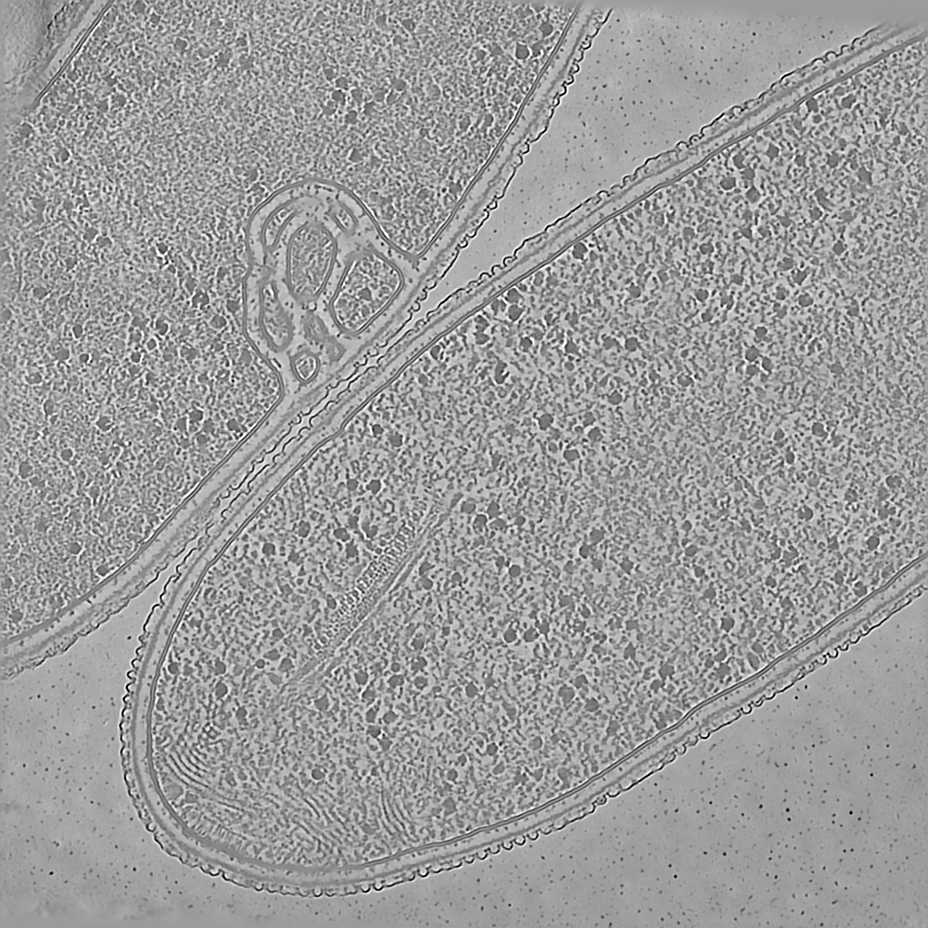
We illustrate Icecream on our favorite tomogram: T. kivui, an anaerobic bacterium that efficiently fixates carbon. The interest of T. kivui is that deep learning methods are known to improve a lot compared to standard filtered back-projection.
In the following figures, we reported Icecream along with:
- DeepDeWedge, conceptually the closest to Icecream. Both implement similar equivariant losses, with the main difference that Icecream treats symmetry in a way consistent with the recent equivariant imaging framework. This amounts to applying the trained network twice at inference.
- CryoLithe, a supervised learning method. The strength of CryoLithe is that reconstructing such a volume takes 4 minutes, against 12 hours for Icecream and 24 hours for DeepDeWedge. A strong or not so strong feature is that CryoLithe doesn’t require any parameters, only the aligned tilt-series.
- Filtered back-projection, or weighted back projection. This was performed using Imod and the
tiltfunction.
About technical details. This is tomo2_L1G1 from EMPIAR-11058. The pixel size is 14.08Å and the tomogram contains 928 x 928 x 464 pixels. Two tomograms were obtained by spliting the tilt angles.