Examples of reconstructions
EMPIAR 11830 - Chlamydomonas reinhardtii prepared using cryo-plasmaFIB milling
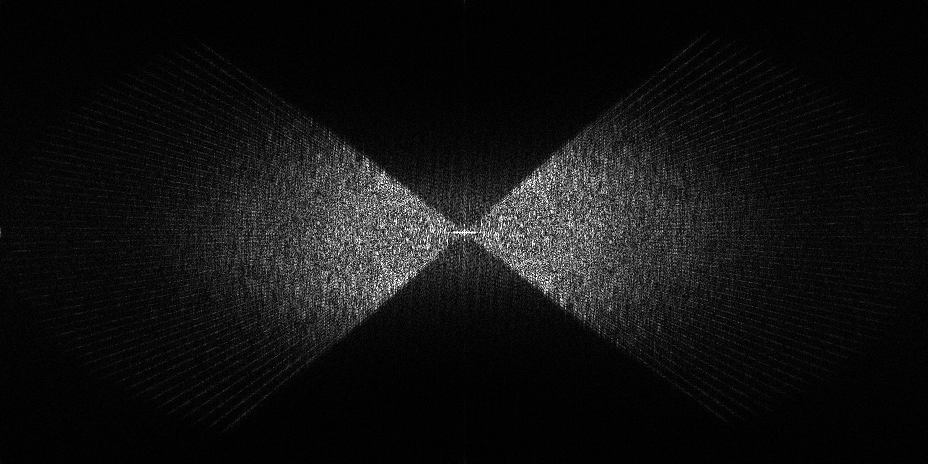
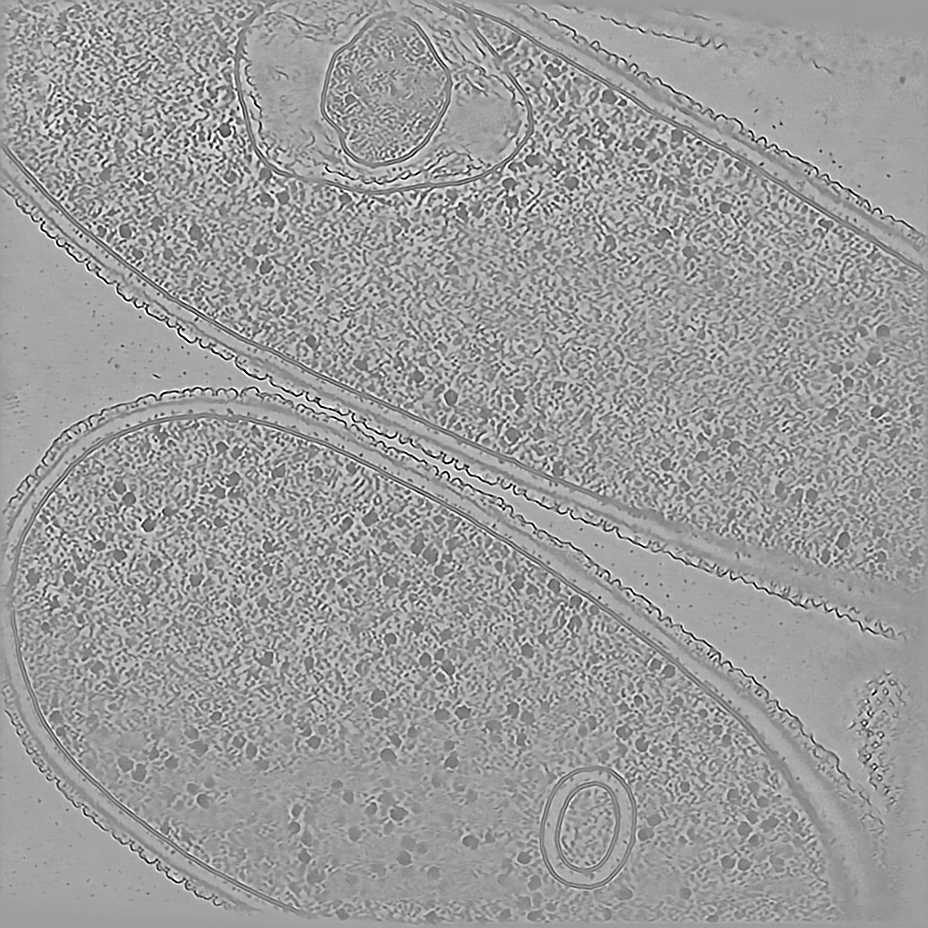
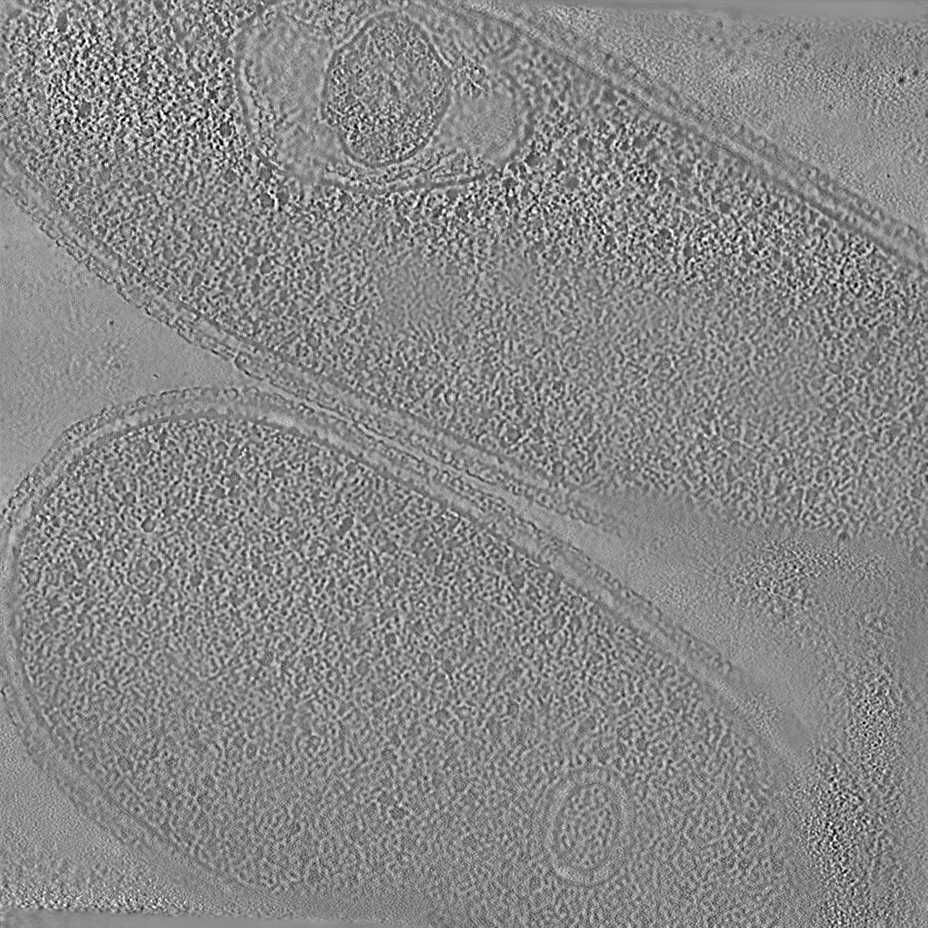
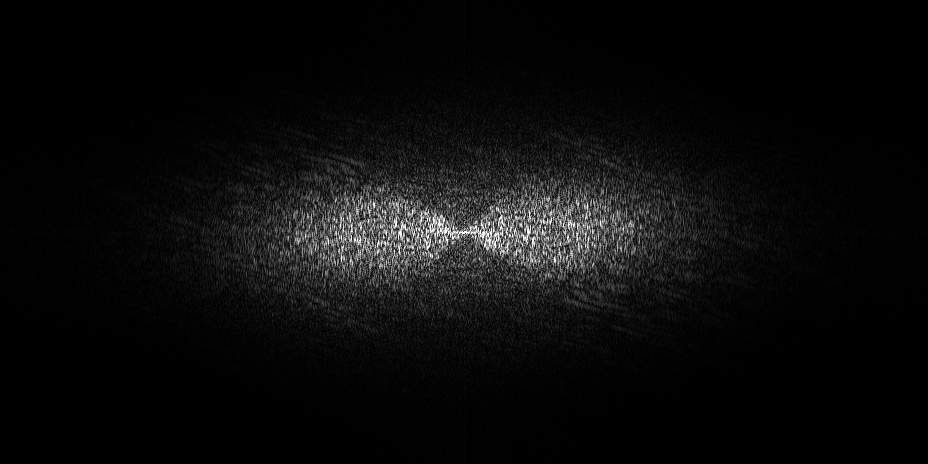
The pixel size is 7.84Å and the tomogram contains 1024 x 1024 x 512 pixels. Two tomograms were obtained by spliting the dose. Below we display both Icecream and DeepDeWedge reconstruction, as well as the stretched amplitude of their Fourier transform on the XZ plane.
Tips: Use the mouse wheel to zoom in and out at specific locations. The buttons below allows to zoom in and out from the central pixel. The slice slider allows you to chose the z-slice to visualize.
Tomogram 9 from EMPIAR-11830
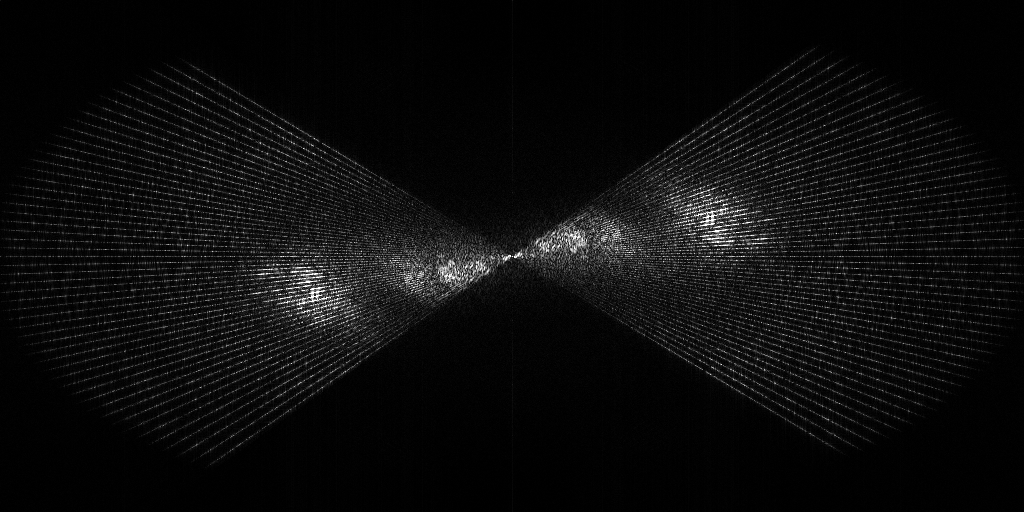
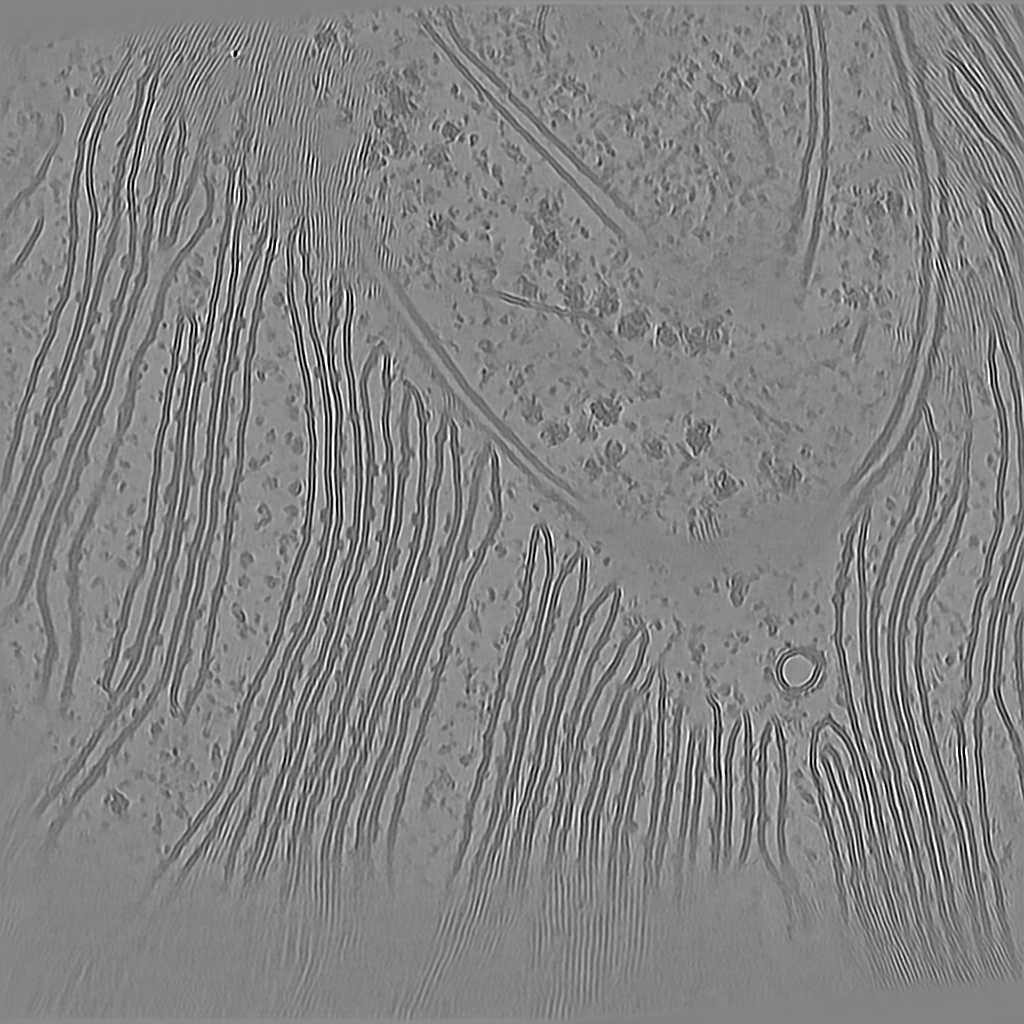
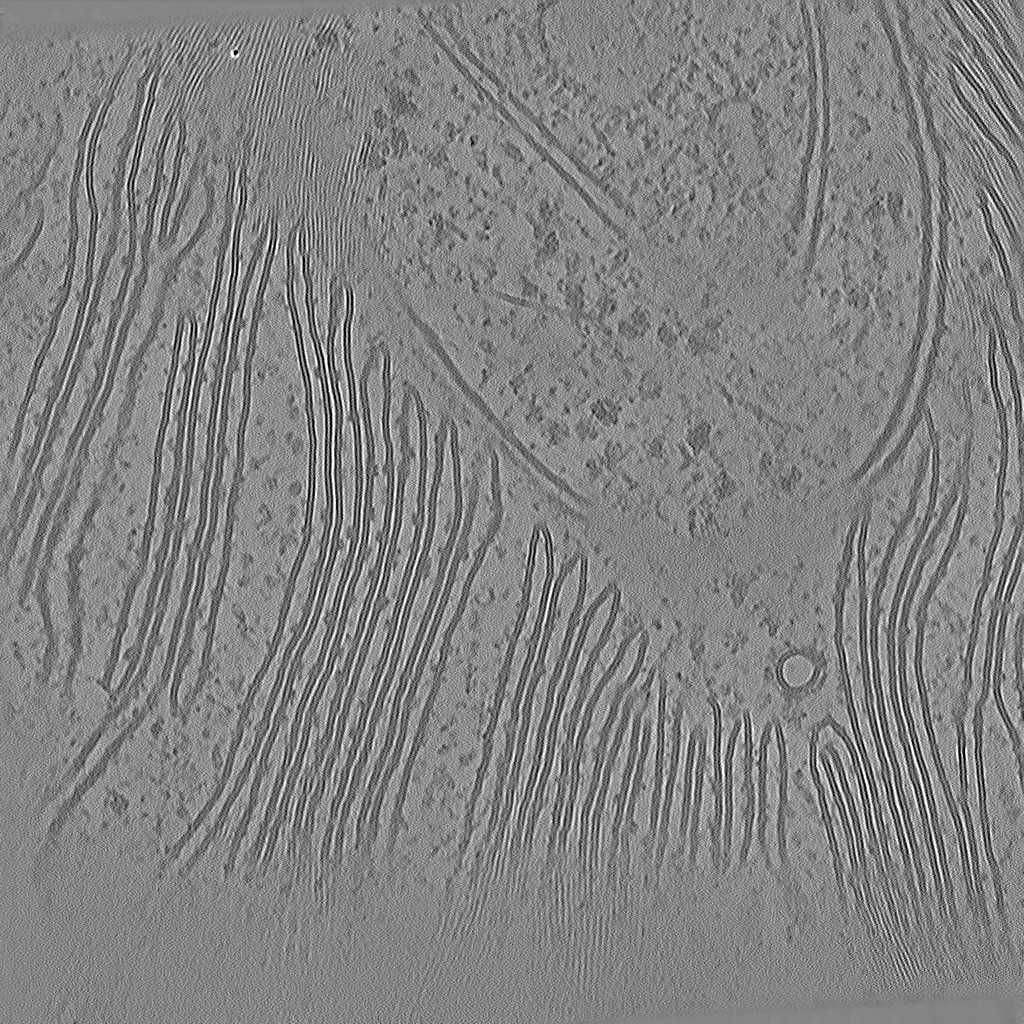


Tomogram 2 from EMPIAR-11830
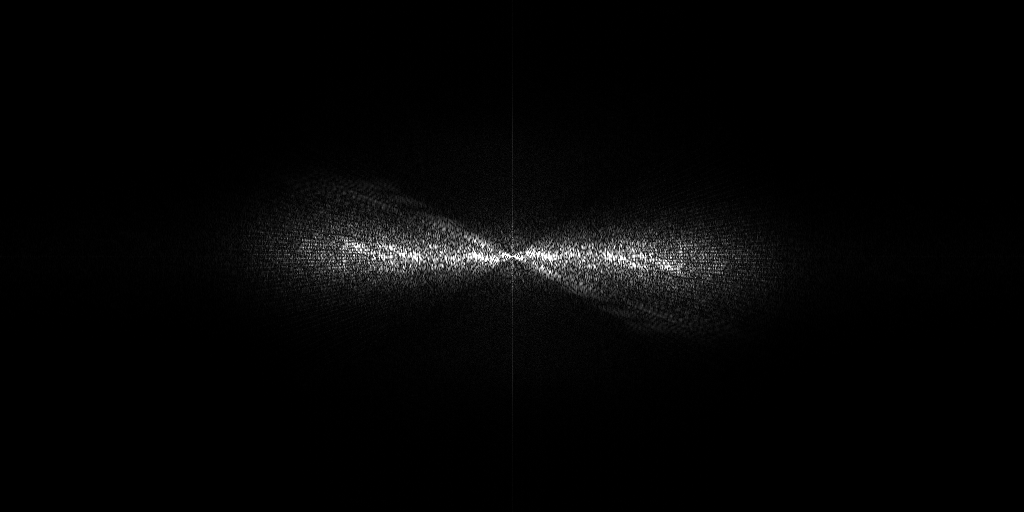
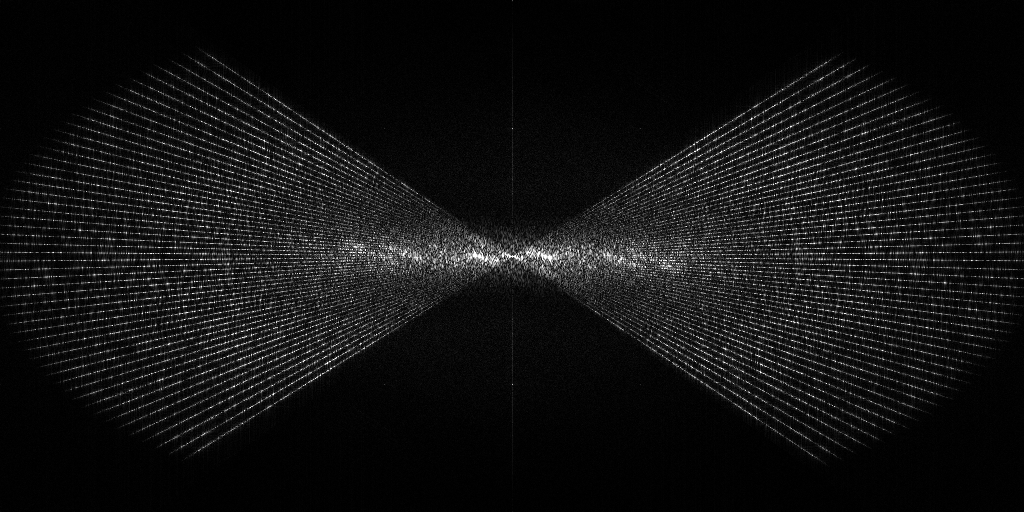
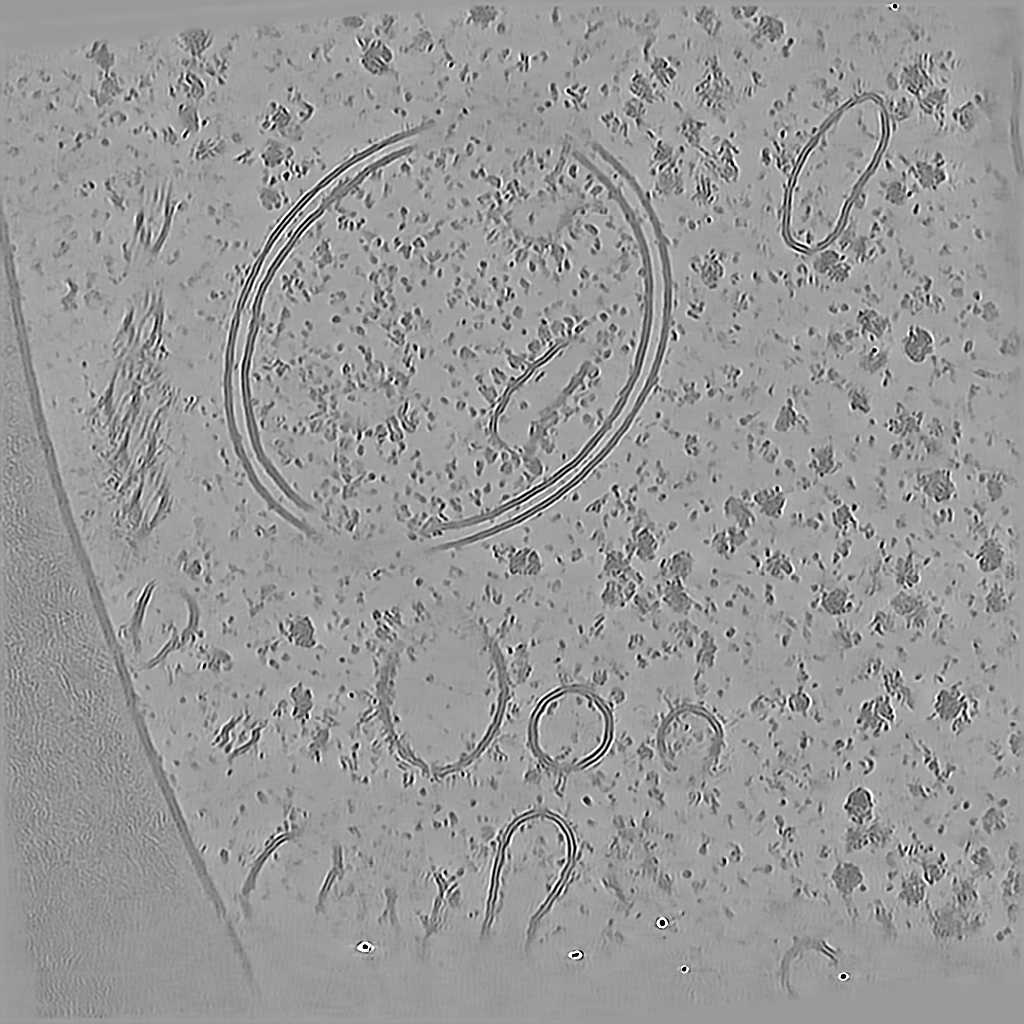
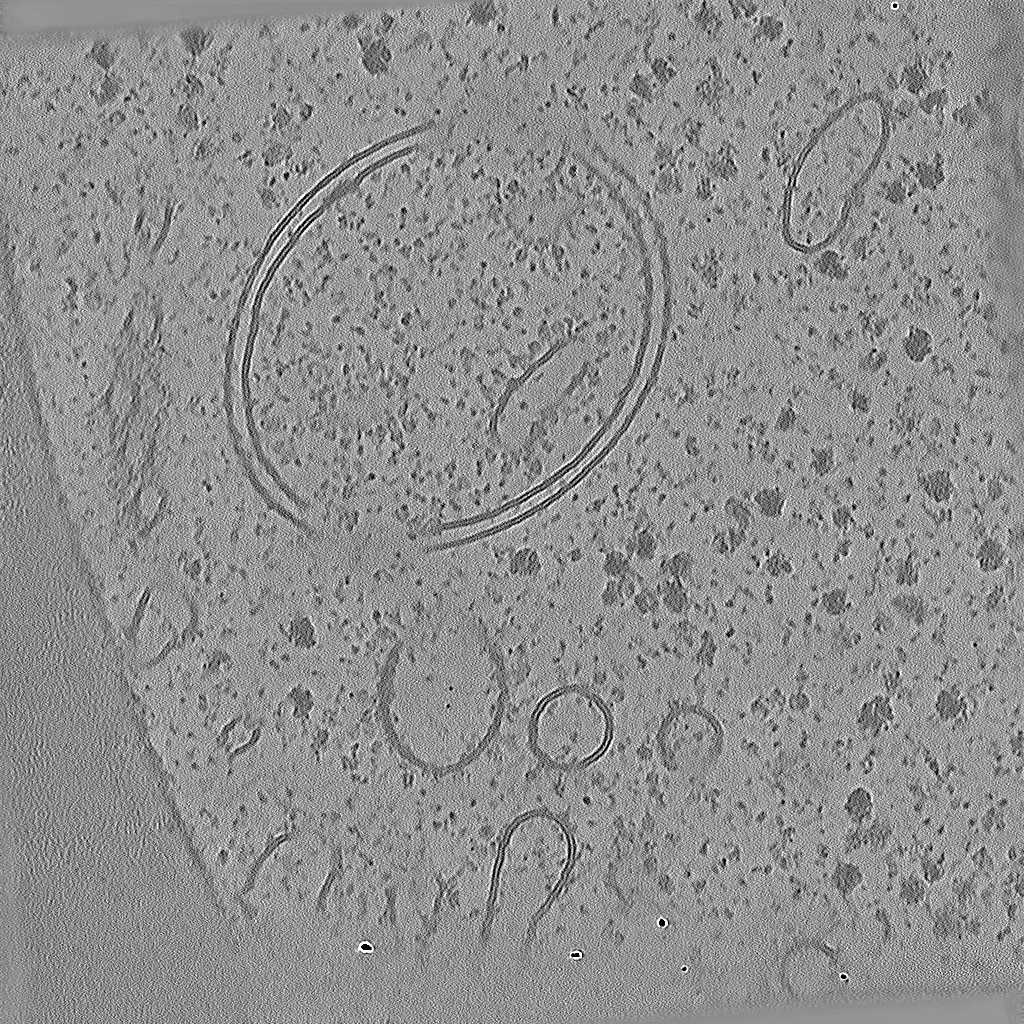


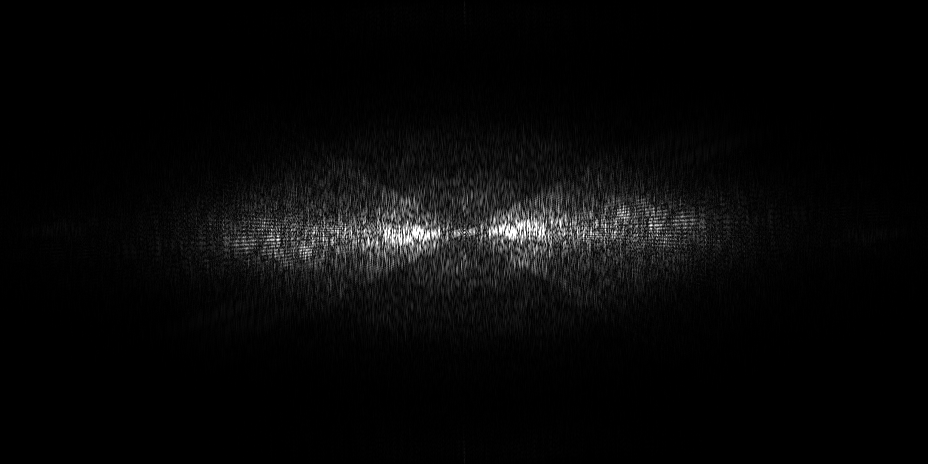
EMPIAR 12612 - Molecular architecture of thylakoid membranes within intact spinach chloroplasts
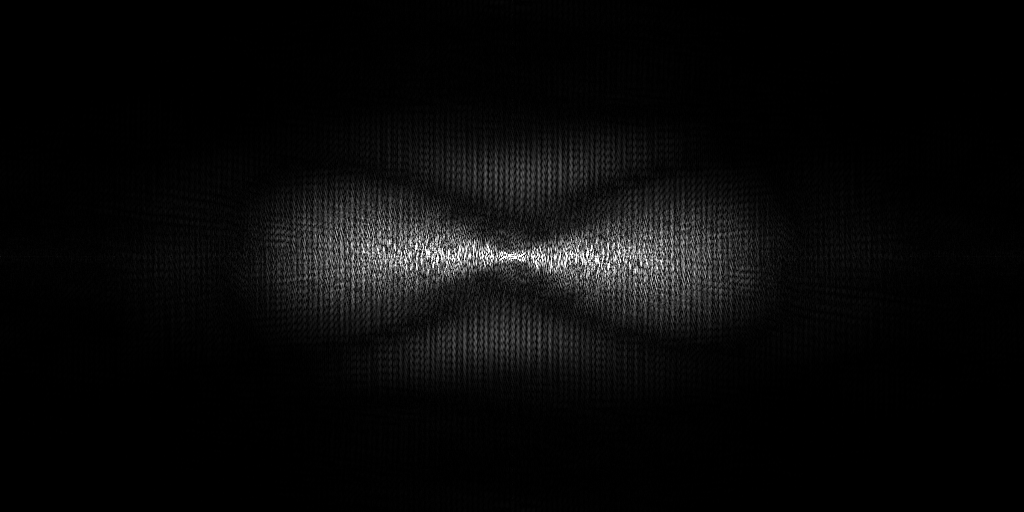
The pixel size is 14.08Å and the tomogram contains 928 x 928 x 464 pixels. Two tomograms were obtained by spliting the dose. This is tomogram 38 from EMPIAR-12612.
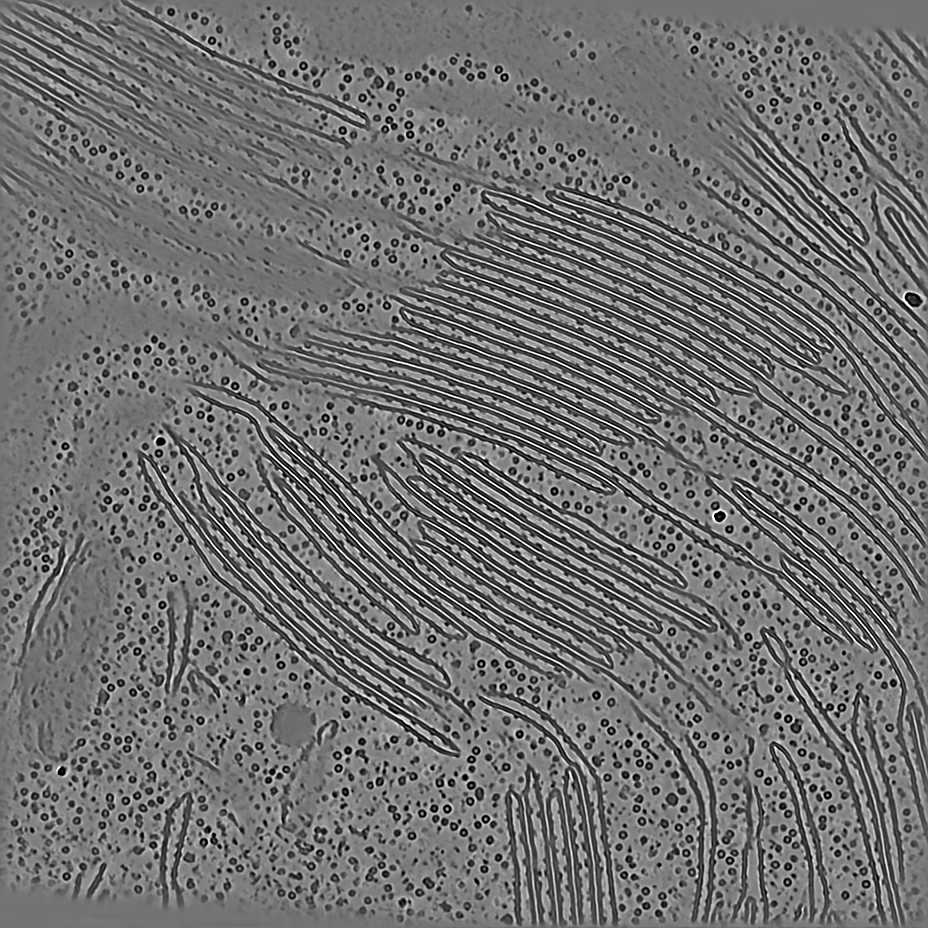
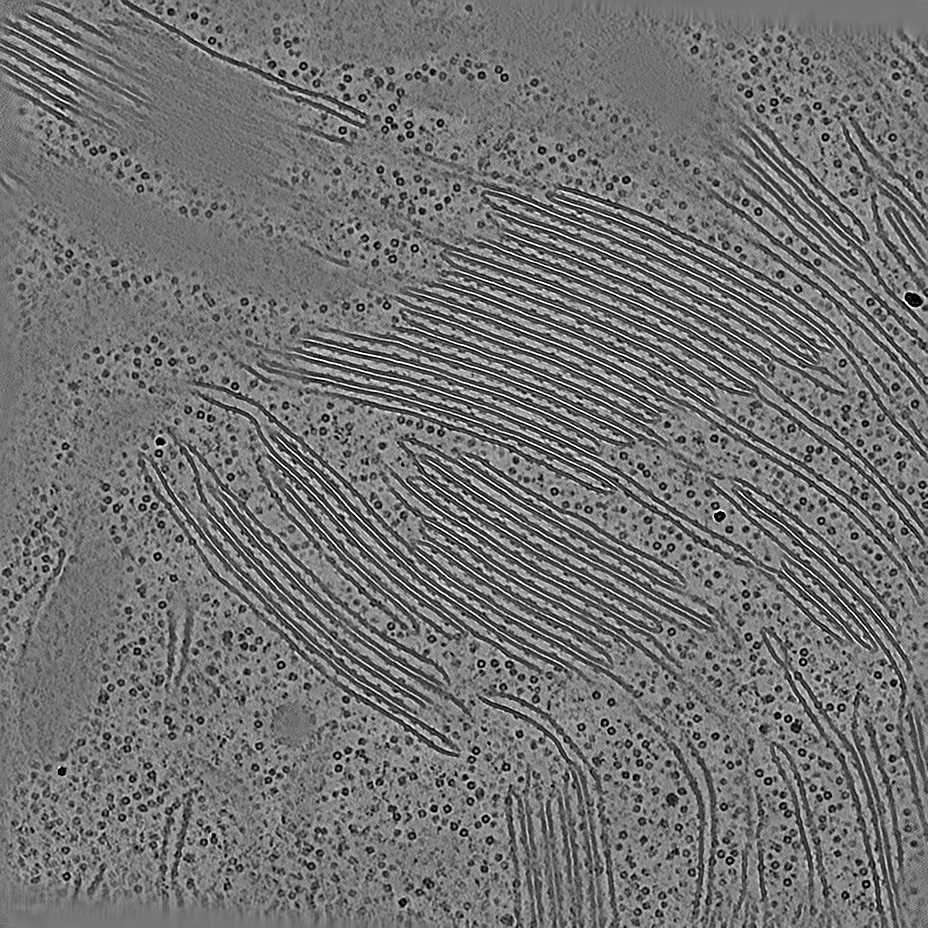
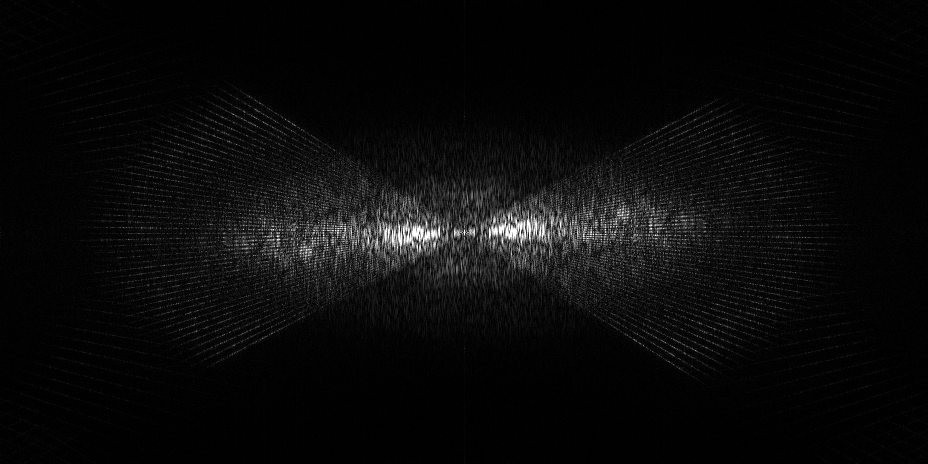



EMPIAR 11538 - HEK293T cells
The pixel size is 4Å and the tomogram contains 1024 x 1024 x 512 pixels. Two tomograms were obtained by spliting the tilt angles. This is tomogram 1435 from EMPIAR-11538.
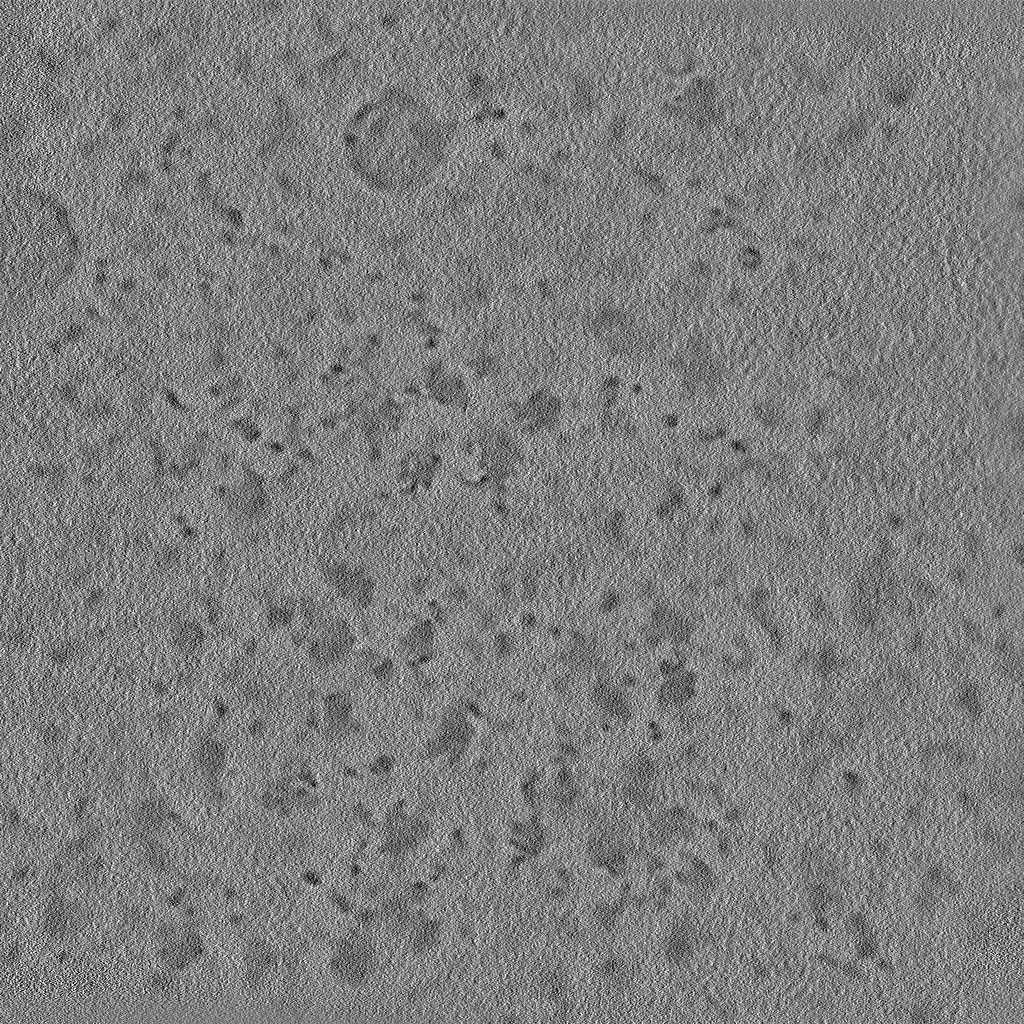
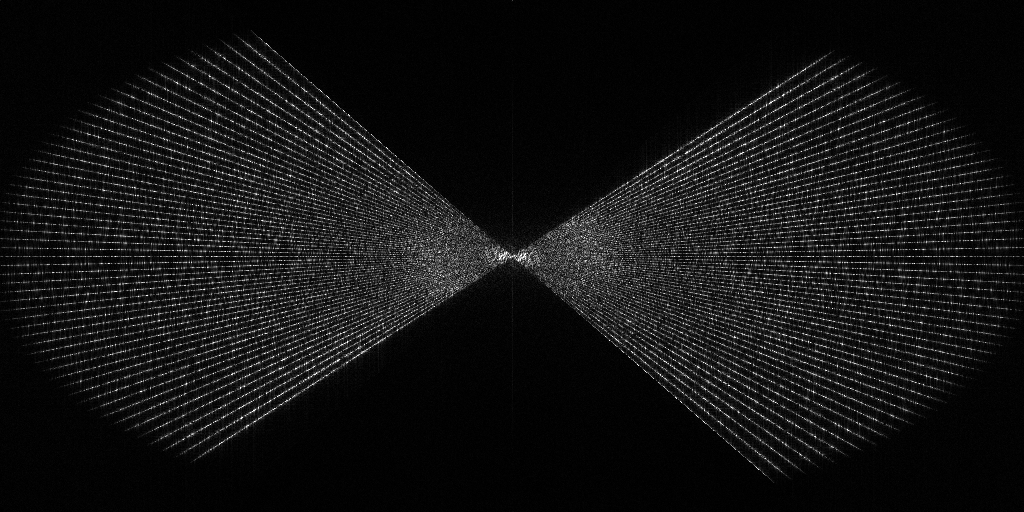
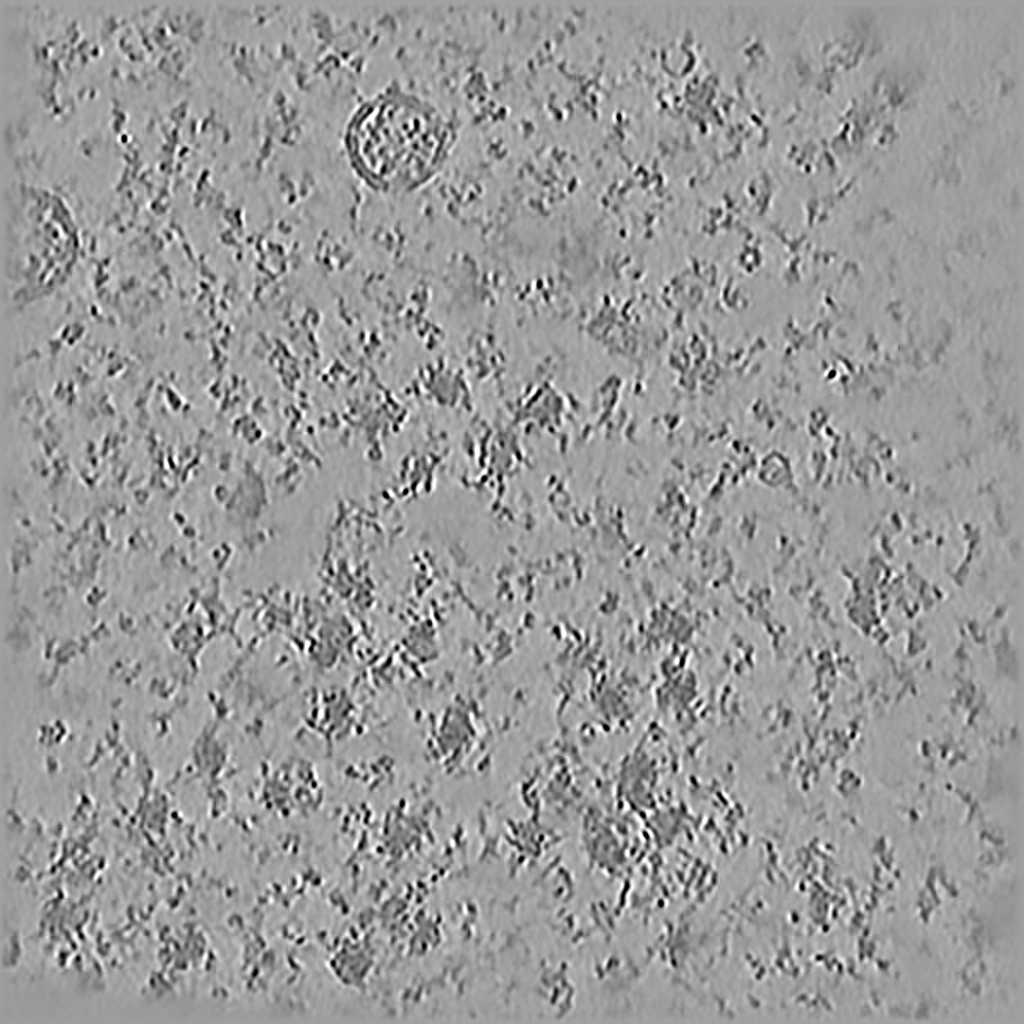


EMPIAR 11658 - S. cerevisiae prepared using cryo-plasmaFIB milling
The pixel size is 7.84Å and the tomogram contains 1024 x 1024 x 512 pixels. Two tomograms were obtained by spliting the tilt angles. This is tomogram 1 from EMPIAR-11658.
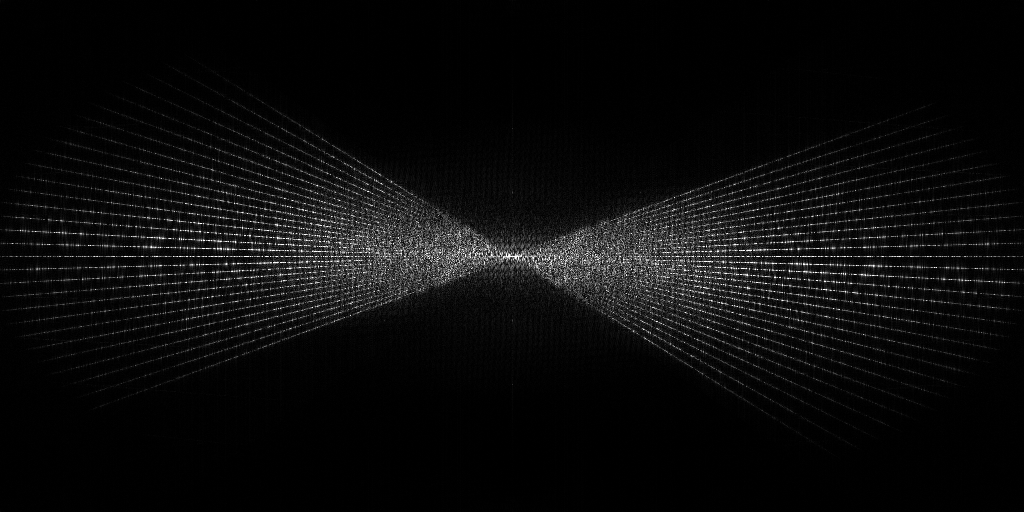
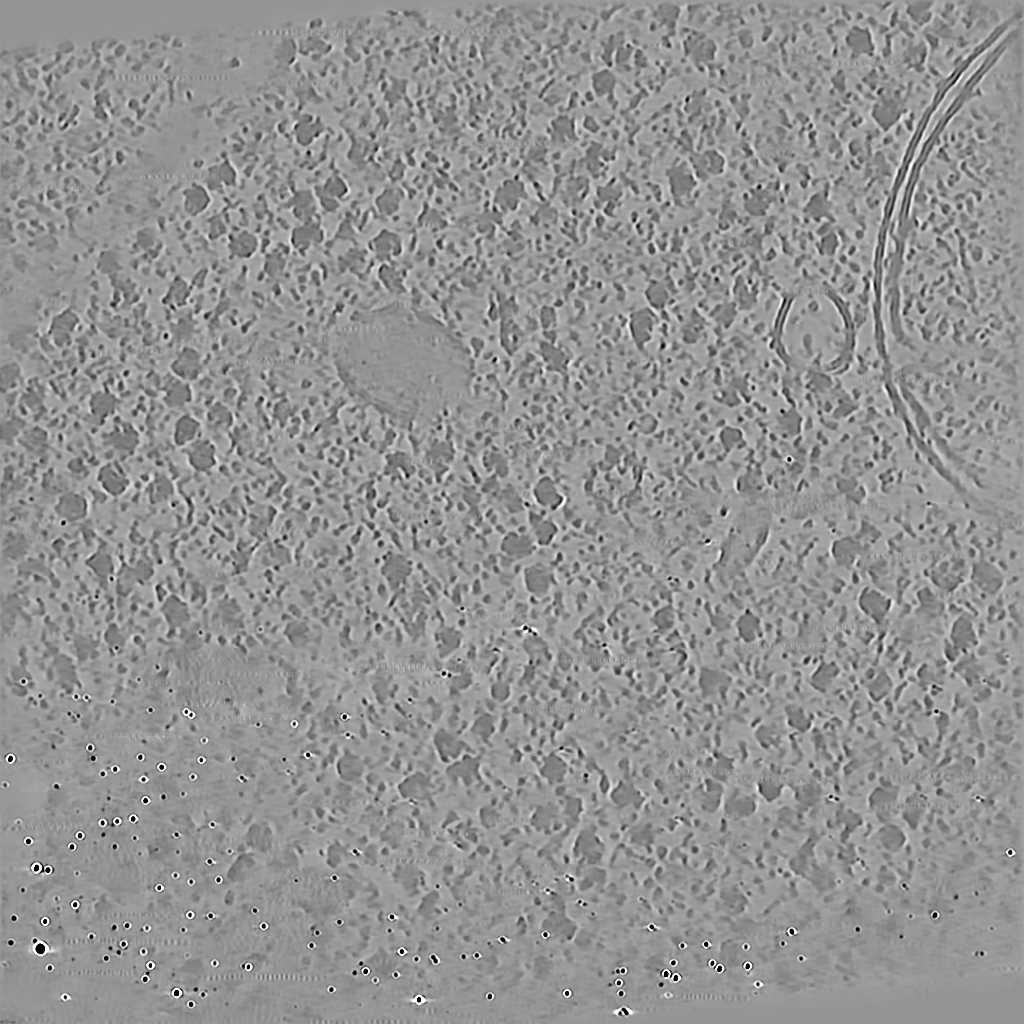
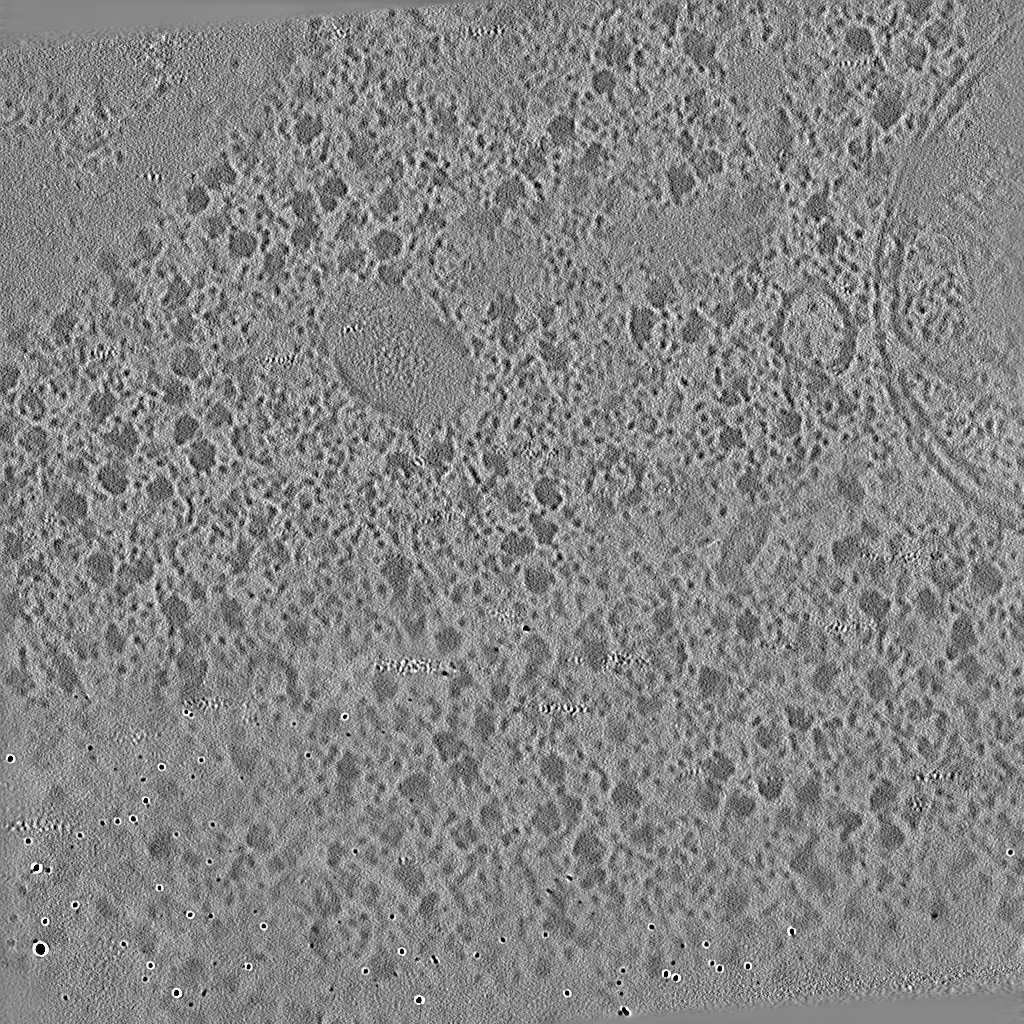


EMPIAR 11058 - T. kivui cells
The pixel size is 14.08Å and the tomogram contains 928 x 928 x 464 pixels. Two tomograms were obtained by spliting the tilt angles. This is tomogram 3 from EMPIAR-11058.